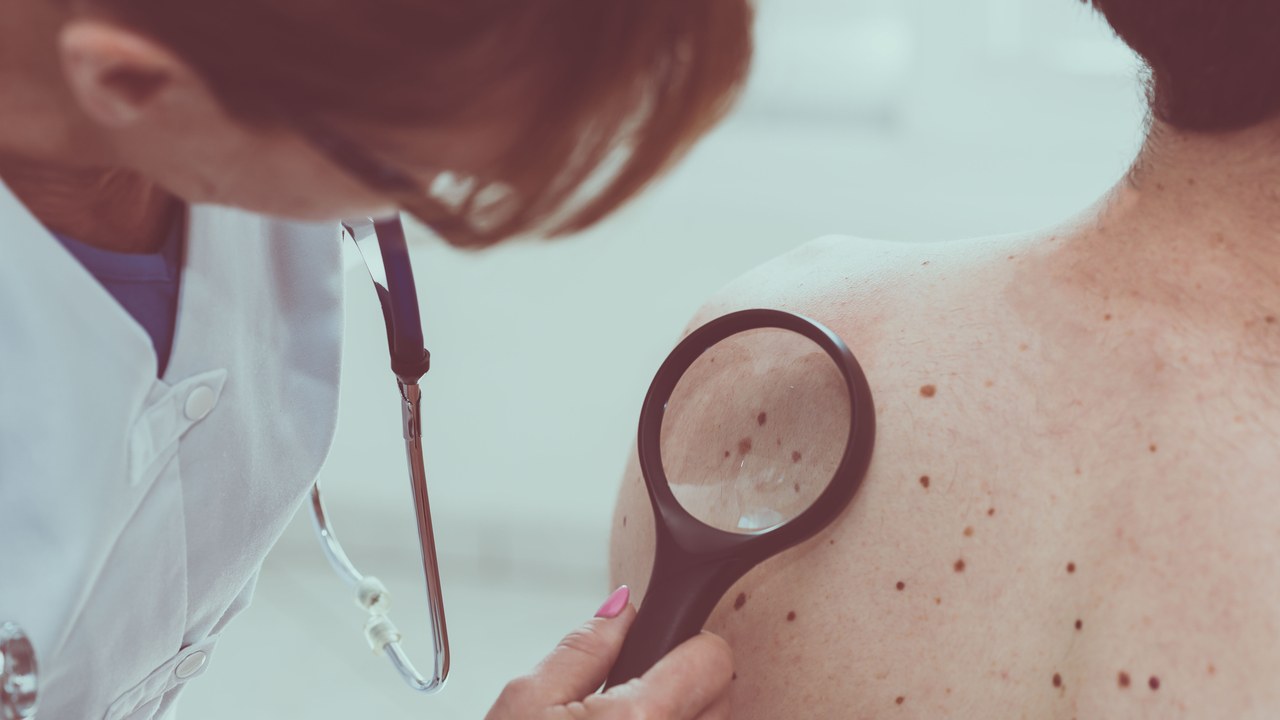
Yes! It is. Friedrich Miescher extracted DNA from the blood first time in 1869. DNA extraction is a technique performed for the purification of DNA with the help of physical or chemical methods from a blood sample by separation of DNA from cell constituents, like cell membranes, proteins, and lipids.
DNA extraction is the basic procedure employed in the molecular laboratory. The procedure may provide efficient extraction if an absolute DNA is used. Quantity of DNA matters as well. DNA should be pure and is devoid of contaminants, including RNA and proteins. Manual procedures are used for DNA extraction but extraction kits are available in market as well. You can use several kinds of the tissues for this purpose including blood, body fluids, formalin-fixed paraffin-embedded tissues, direct Fine needle aspiration cytology (FNAC) aspirate, frozen tissue section, etc.,
What are the techniques to extract DNA?
DNA extraction consists of lysis (breakdown) of the cells and solubilizing DNA (to stable pH), which is further followed by enzymatic or chemical methods to take out macromolecules, proteins, lipids, and RNA. Following techniques are included in DNA extraction:
- Organic extraction (PCI- phenol chloroform isoamyl method)
- Inorganic technique (salting out and proteinase K treatment)
- Adsorption method (silica–gel membrane)
Let’s discuss the first one organic extraction with PCI suspension. This technique is labor intensive and time consuming but yields efficient results.
Collection of blood sample
First step is to collect the blood sample. Collect 5mL of blood aseptically from jugular vein of any specie (horse in this case) in 15mL falcon tube. 25 microlitre anticoagulant 0.5 M EDTA (ethylene diamide tetra-acetic acid) was already present in falcon tube.
Storage of blood sample
In order to prevent any contamination, blood sample was stored very after its collection. Place the blood sample on ice at 0°C for 24 hours in your laboratory freezer.
Procedure and technicalities
Primarily, you have to thaw the falcon tube containing blood sample by putting it at room temperature. Now add 500 microliter of washing buffer in 500 microliter of blood. Washing buffer helps disrupting red blood cells, and therefore it is also called as lysis buffer. Mix it thoroughly. Then, centrifuge the sample at 14,500 rpm for 15 minutes at 4°C for efficient results. As a result of centrifugation, pellet will form at the bottom of tube. Discard the supernatant and break the pellet gently by taping on the tube. Washing buffer will be added again. Continue repeating procedure by adding washing buffer until pellet color becomes dull white or pure white.

Add buffer 10 about 300 microliter, proteinase K about 50 microliter (after thawing it), and SDS about 90 microliter in 200microliter of blood. Mix these contents on vortex for 10 second and incubate it at 37°C overnight in a shaking incubator. Add Phenol Chloroform Isoamyl Alcohol (PCI) suspension in equal amount into the mixture present in 15 mL microcentrifuge eppendorf. Mix it altogether and centrifuge at 14,500 rpm for 20 minutes.

Formation of three layers
As a result of centrifuge, three layers are formed with the components including:
- Upper aqueous layer containing DNA
- In between, there exists a white layer amalgamated with high protein concentration
- Lower layer containing phenol
Transfer the upper layer containing DNA into microcentrifuge tube. Absolute chilled isopropanol is added in equal amount of supernatant to condense the DNA. To observe DNA fragments, invert the tube till small threads of DNA starts appearing.
If you have seen DNA structures, this is an indication DNA has been extracted from your blood sample. Now, place the tubes at room temperature for almost 10 minutes. Then, pellet is washed with 100 microliter of 70% ethanol on the shaking stand. Centrifuge the mixture again at 4000 rpm at 0°C.
Heat shock to DNA
Expose the chemical mixture to sudden heat, a pressure difference between inside and outside of the cell is created as a result. This helps maintaining the constant conditions when contact the mixture immediately with ice. Primarily, this is used to remove all possible contaminants that could be present in the mixture. First of all, 100 microliter water from injections is added into the dried DNA pellet. Mix it thoroughly by placing eppendorf on vortex mixer. It results in alteration in consistency, thus, you can observe a clear mixture. Place the eppendorf on hot water bath at 50°C to 70°C for at least 30 minutes. Very after the hot water bath, put the eppendorf into ice, deeply for absolute cold pressure for no more than 10 minutes. Now, at last, store the tube at room temperature.
Reagents used in DNA extraction and their functions
Reagents are chemical compounds added to any mixture to cause a chemical reaction. Organic method of DNA extraction uses large number of chemical ingredients. Let’s talk about these reagents and their functions:
1. EDTA
Ethylene diamide tetra-acetic acid (EDTA) helps binding certain metal ions. It prevents blood sample from clotting leading to better results.
2. Washing buffer or lysis buffer
It contains high concentration of ethanol and isopropanol and plays an important role in maintaining pH. Washing buffer solubilizes all components effectively and removes the protein traces present in blood. It inhibits RNase free water as well in order to avoid degrading of RNA.
3. Buffer 10
Buffer 10 keeps the pH stable over cell lysis and extraction. It is responsible for the protection of DNA.
4. Proteinase K
Self-digested proteinase K (20mg/mL) contains proteinase K pack of 100 mg and tris-EDTA-SDS (100 mM Tris, pH 8.0, 40 mM EDTA, 0.05% SDS) 5 mL. The most important role of proteinase K is to inactivate the RNase. It denatures the protein present in blood sample and release nucleic acids.
5. SDS
10% Sodium Dedocyl Sulfate (SDS) consists of Sodium Dedocyl Sulfate (Sodium Lauryl Sulfate) of 10 gm and deionized water 100 mL. It helps in cell lysis and denaturing of the protein structures as it imparts negative charge to proteins. SDS also breaks the lipid bilayer of white blood cells.
6. Isopropanol
Isopropanol works by precipitating DNA because nucleic acids are insoluble in ethanol, that’s why to ensure the process of accurate precipitation isopropanol is added. 75% ethanol (absolute ethanol 70 mL and deionized water 30 mL) is not suitable for precipitation, so it is better to mix and wash away salts while leaving behind most of the DNA or RNA. Isopropanol is also used in washing or storing of DNA.
7. Ethanol
Ethanol is added to remove some (ideally all) salts from the pellet. It assists precipitating DNA out of aqueous solution and promotes DNA aggregation. Ethanol diminishes the solvation shell surrounding the DNA, therefore letting the DNA to precipitate in pellet form.